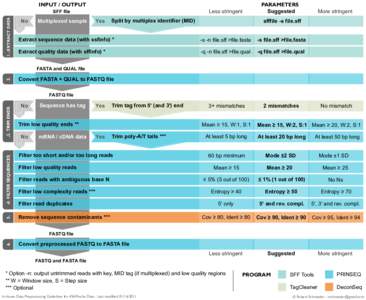
Date: 2013-02-11 17:31:18Molecular biology Gene expression Biological databases Protein methods Chip-sequencing RNA-Seq FASTA format TRANSFAC FASTQ format Biology Bioinformatics Biochemistry | |  Workflow for ChIP-seq & RNA-seq data analysis, GRS 2012 4 WORKFLOW FOR ANALYSIS OF CHIP-SEQ DATA INCLUDING ANALYSIS OF ENRICHED TRANCRIPTION FACTOR BINDING SITES -Gavin Schnitzler[removed]QUALITY CONTROL ON SEQUE Workflow for ChIP-seq & RNA-seq data analysis, GRS 2012 4 WORKFLOW FOR ANALYSIS OF CHIP-SEQ DATA INCLUDING ANALYSIS OF ENRICHED TRANCRIPTION FACTOR BINDING SITES -Gavin Schnitzler[removed]QUALITY CONTROL ON SEQUE
Add to Reading ListSource URL: sites.tufts.eduDownload Document from Source Website File Size: 162,50 KBShare Document on Facebook
|
 Workflow for ChIP-seq & RNA-seq data analysis, GRS 2012 4 WORKFLOW FOR ANALYSIS OF CHIP-SEQ DATA INCLUDING ANALYSIS OF ENRICHED TRANCRIPTION FACTOR BINDING SITES -Gavin Schnitzler[removed]QUALITY CONTROL ON SEQUE
Workflow for ChIP-seq & RNA-seq data analysis, GRS 2012 4 WORKFLOW FOR ANALYSIS OF CHIP-SEQ DATA INCLUDING ANALYSIS OF ENRICHED TRANCRIPTION FACTOR BINDING SITES -Gavin Schnitzler[removed]QUALITY CONTROL ON SEQUE